


Five Molecular Structures Solved in 24 Hours
by Lisa Horanyi and Carole VanSickle

Zhi-Jie Liu’s team just set a new world record for speed. But they’re not Olympic runners or NASCAR drivers. They’re molecular biologists at the University of Georgia who determined the structures of five proteins in less than 24 hours.
“Nothing like this has ever happened before,” said Liu upon his return this past May from the U.S. Department of Energy’s Argonne National Laboratory Advanced Photon Source, the site of their record-breaking performance. APS, which is run by the University of Chicago and located about 25 miles west of its campus, produces some of the fastest and most brilliant X-ray beams, or beamlines, in the world. When focused on crystallized proteins, beamlines help scientists illuminate both the proteins’ structures and functions.
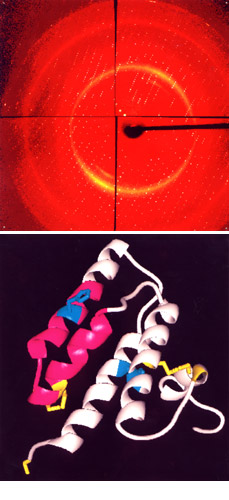
To examine the molecular structures of proteins, scientists grow proteins in crystalline form. Once a crystal has been selected, it is placed in the beam of the X-ray and rotated in very small increments. A new X-ray pattern is collected for each increment, and these patterns can be “sewn” together to virtually reconstruct the protein.
Liu said that the record-breaking achievement is the culmination of advancements made to the Southeast Regional Collaborative Access Team’s (SER-CAT’s) beamlines, which are among the most intense and stable in the country, and the Southeast Collaboratory for Structural Genomics, headquartered at UGA.
Liu’s team combined technique with technology, including “direct crystallography,” which was proposed by Bi-Cheng Wang, the Ramsey-Georgia Research Alliance Eminent Scholar in structural biology at UGA and director of Liu’s lab. Direct crystallography employs a multidisciplinary team approach that goes straight to the determination of structures, eliminating much time from the procedure. Liu leads the newly formed crystallomics group, which works to unify all aspects of protein-crystal production instead of relying on isolated steps performed in different labs.
Prior to the spring 2005 visit to SER-CAT, the team’s best performance ever achieved with protein crystallography was three structures in 24 hours. Wang credits not only Liu’s team but also the APS facility and newly refined equipment. “We have increased the stability of the beam and nearly eliminated the wobble that was there before,” he said.
According to Liu, the team plans to publish their new data- collection technique, which will help many other crystallographers.
“A state-of-the-art robot picked the best crystals ahead of time to give the researchers a good idea of what they could expect to find when they placed the crystals in the beamline,” he said. “Most crystallographers run a job then check results. Our program is like having hundreds of crystallographers working together.”
For more information on SER-CAT, access http://www.ser-cat.org and on the Southeast Collaboratory for Structural Genomics, access http://www.secsg.org or contact Zhi-Jie Liu at xtalfarmer@gmail.com.
For comments or for information please e-mail: rcomm@uga.edu
To contact the webmaster please email: ovprweb@uga.edu
![]()